Two-component
system (TCS) signal transduction pathways, are found
across all three domains of life allowing adaptive responses to changes in
environmental conditions. However, they are mainly found in bacteria where they
control multiple aspects of bacterial metabolism, such as cell differentiation
and morphogenesis, central metabolism, motility, biofilm formation and
virulence. These systems were classically described as the association of two
proteins that communicate through a His-Asp phosphorelay.
Typical TCSs comprise a histidine kinase sensor protein which is capable of autophosphorylation
on a conserved His residue, before transferring the phosphoryl
group to a conserved Asp residue within the receiver domain (REC) of a response
regulator.
The large number
of TCS protein sequences available demands user-friendly databases to facilitate inter-genomic and
intra-genomic analyses. We
have therefore developed a novel resource, the P2CS (Prokaryotic 2-Component Systems) database, that contains the TCSs of all available bacterial and archaeal genomes, and 39 metagenomes. Our objective was to
provide an easy to use environment for validation by experts, according
to their fields/organisms of interest, with the data being completely available and
consultable by all of the scientific community.
From
the P2CS website the user can:
Browse
genome and metagenome TCS predictions and manually curated proteins
Search
for a sequence id or domain family
The
P2CS homepage contains a navigation bar that allows database browsing. Among
the menus, users will also find P2CS Browse, which links directly to sortable
lists of analysed genomes, plasmids and metagenomes.
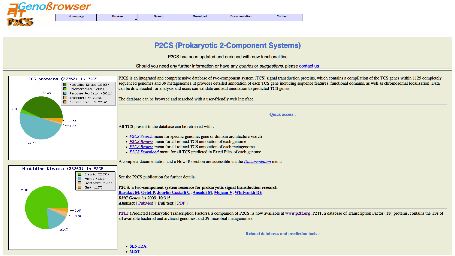
The P2CS database
contains the TCSs of all available bacterial and archaeal genomes, and 39 metagenomes.
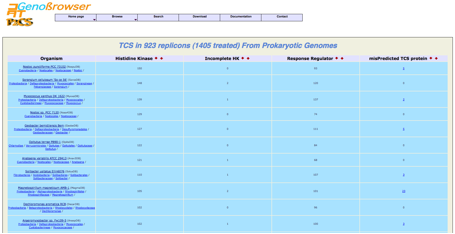
The
selection of a microbe or microbiome displays the
result of the P2CS analysis process.
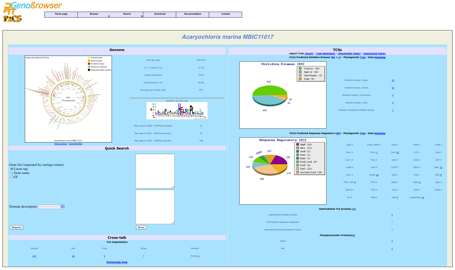
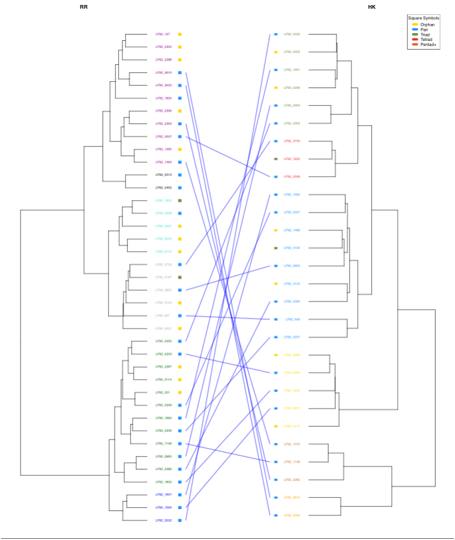
The genome page contains clickable links to RR and HK trees,
heatmaps and double dendrograms
displaying TCS proteins associations (replicons with more than 10 RRs and 10 HKs).
These pages include several options such as zooming in or out.

The genome page
shows also global counts of
the different categories of
TCSs and detailed class counts of each category. Each class result provides a clickable link to a detailed gene list.
It shows also a
search module, based
on :
·
locus-tag
·
gene
name
·
gi number
·
domain
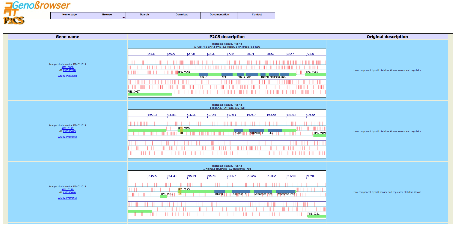
Selecting an object from the list identifiers
displays a detailed gene
description page with an image representing
the gene in the appropriate
frame. Red vertical lines represent stop codons and green lines
represent potential start codons.
Blast searches can be performed
for the sequence, using external links to numerous databases.
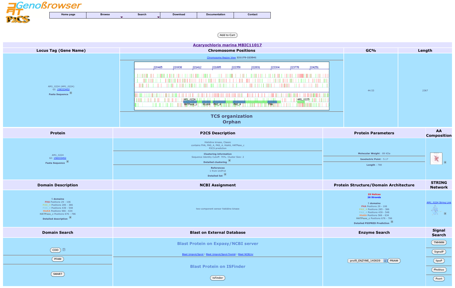
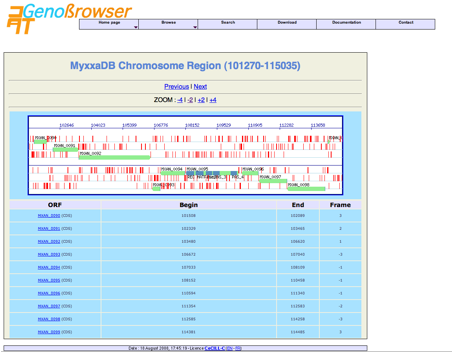
The gene
description page contains a link to a cartographic gene context (Chromosome
Region View), with several options such as zooming in or out, moving along the
chromosome, displaying genes in upstream or downstream regions and drawing
genes.
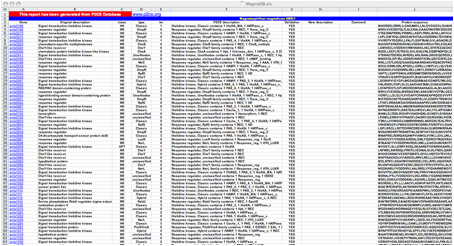
P2CS was designed to allow download of TCS data in tab-delimited
format and generates a file compatible with spreadsheet programs such as Excel. Users can also download
for each genome and metagenome a multi-Fasta file (nucleotide or protein sequences).
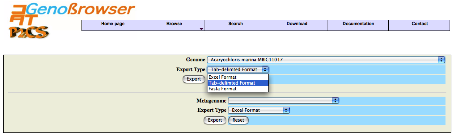
An example of a
Excel file output.